Optimization with APOSMM¶
This tutorial demonstrates libEnsemble’s capability to identify multiple minima
of simulation output using the built-in APOSMM
(Asynchronously Parallel Optimization Solver for finding Multiple Minima)
gen_f. In this tutorial, we’ll create a simple
simulation sim_f that defines a function with
multiple minima, then write a libEnsemble calling script that imports APOSMM and
parameterizes it to check for minima over a domain of outputs from our sim_f.
Six-Hump Camel Simulation Function¶
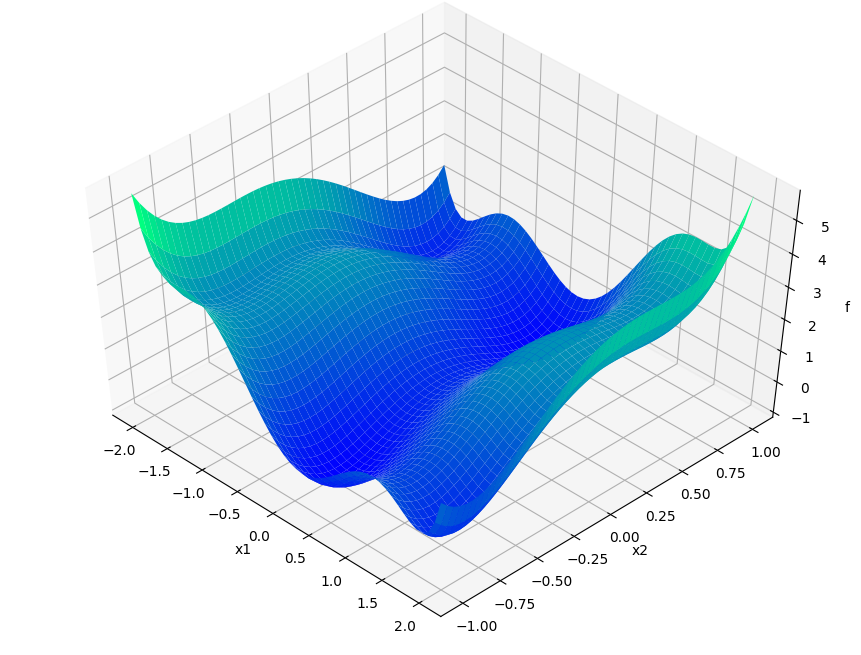
Describing APOSMM’s operations is simpler with a given function on which to depict evaluations. We’ll use the Six-Hump Camel function, known to have six global minima. A sample space of this function, containing all minima, appears below:
Create a new Python file named six_hump_camel.py. This will be our
simulator callable, incorporating the above function. Write the following:
1def six_hump_camel_func(x):
2 """Six-Hump Camel function definition"""
3 x1 = x["x1"]
4 x2 = x["x2"]
5 term1 = (4 - 2.1 * x1**2 + (x1**4) / 3) * x1**2
6 term2 = x1 * x2
7 term3 = (-4 + 4 * x2**2) * x2**2
8
9 return {"f": term1 + term2 + term3}
APOSMM Operations¶
APOSMM coordinates multiple local optimization runs starting from a collection of sample points. These local optimization runs occur concurrently, and can incorporate a variety of optimization methods, including from NLopt, PETSc/TAO, SciPy, or other external scripts.
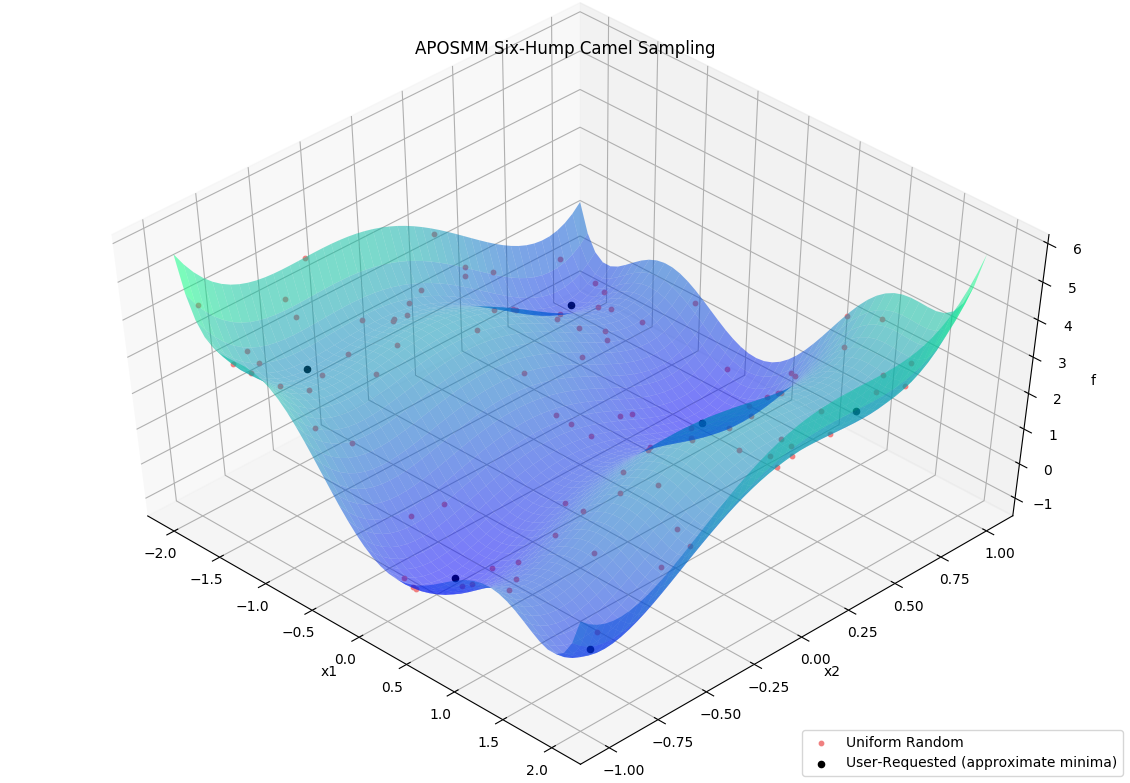
Before APOSMM can start local optimization runs, some number of uniformly sampled points must be evaluated (if no prior simulation evaluations are provided). User-requested sample points can also be provided to APOSMM:
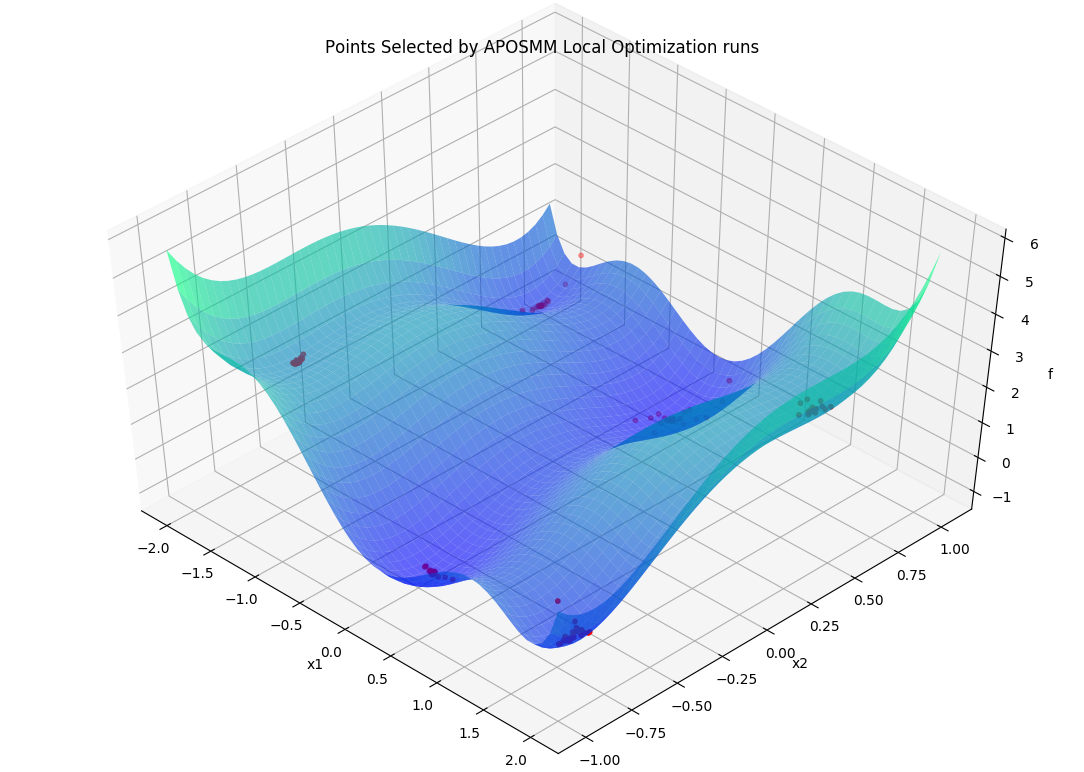
Specifically, APOSMM will begin local optimization runs from evaluated points that
don’t have points with smaller function values nearby (within a threshold
r_k). For the above example, after APOSMM receives the evaluations of the
uniformly sampled points, it will begin at most max_active_runs local
optimization runs.
As function values are returned to APOSMM, APOSMM gives them to the
corresponding local optimization runs so they can generate the next point(s) in
their runs; such points are then returned by APOSMM to the manager to be
evaluated by the simulation routine. As runs complete (i.e., a minimum is
found, or some termination criteria for the local optimization run is
satisfied), additional local optimization runs may be started or additional
uniformly sampled points may be evaluated. This continues until a STOP_TAG
is sent by the manager, for example when the budget of simulation evaluations
has been exhausted, or when a sufficiently “good” simulation output has been
observed.
Throughout, generated and evaluated points are appended to the
History array, with the field
"local_pt" being True if the point is part of a local optimization run,
and "local_min" being True if the point has been ruled a local minimum.
Calling Script¶
Create a new Python file named my_first_aposmm.py. Start by importing
libEnsemble classes, APOSMM, and our simulator callable:
1from six_hump_camel import six_hump_camel_func
2
3import libensemble.gen_funcs
4
5libensemble.gen_funcs.rc.aposmm_optimizers = "scipy"
6
7from libensemble import Ensemble
8from libensemble.gen_classes import APOSMM
9from gest_api.vocs import VOCS
10from libensemble.specs import SimSpecs, GenSpecs, ExitCriteria
APOSMM supports a wide variety of external optimizers. The rc.aposmm_optimizers
statement above indicates to APOSMM which optimization method package to use,
helping prevent unnecessary imports or package installations.
Next, initialize the Ensemble and define our variables and objectives using
a VOCS object:
1if __name__ == "__main__":
2 workflow = Ensemble(parse_args=True)
3
4 vocs = VOCS(
5 variables={"x1": [-2, 2], "x2": [-1, 1], "x1_on_cube": [-2, 2], "x2_on_cube": [-1, 1]},
6 objectives={"f": "MINIMIZE"},
7 )
Notice the addition of x1_on_cube and x2_on_cube. APOSMM requires variables scaled to the unit cube internally. By defining both sets of variables, APOSMM can translate between our actual domain and its internal domain.
Now, configure APOSMM. Because APOSMM internally uses variables named x, x_on_cube, and an objective named f, we must map our VOCS fields to these internal names using variables_mapping:
1 aposmm = APOSMM(
2 vocs,
3 max_active_runs=workflow.nworkers,
4 variables_mapping={"x": ["x1", "x2"], "x_on_cube": ["x1_on_cube", "x2_on_cube"], "f": ["f"]},
5 initial_sample_size=100,
6 localopt_method="scipy_Nelder-Mead",
7 opt_return_codes=[0],
8 )
9
10 workflow.gen_specs = GenSpecs(
11 generator=aposmm,
12 vocs=vocs,
13 batch_size=5,
14 initial_batch_size=10,
15 )
APOSMM is instantiated directly as a standardized generator. It handles its own required fields, simplifying our configurations. opt_return_codes is a list of integers that local optimization methods return when a minimum is detected. SciPy’s Nelder-Mead returns 0.
Finally, we configure the simulation function, exit criteria, and run the workflow. We can also print out any points that APOSMM identified as local minima:
1 workflow.sim_specs = SimSpecs(simulator=six_hump_camel_func, vocs=vocs)
2 workflow.exit_criteria = ExitCriteria(sim_max=2000)
3
4 H, _, _ = workflow.run()
5
6 if workflow.is_manager:
7 # We can map our variables back to an array for easy printing
8 minima = [[row["x1"], row["x2"]] for row in H if row["local_min"]]
9 print("Minima:", minima)
Final Setup, Run, and Output¶
If you haven’t already, install SciPy so APOSMM can access the required optimization method:
pip install scipy
Finally, run this libEnsemble / APOSMM optimization routine with the following:
python my_first_aposmm.py --nworkers 4
Please note that one worker will be “persistent” for APOSMM for the duration of the routine.
After a couple seconds, the output should resemble the following:
Minima: [[0.08988580227184285, -0.7126604246830723], [-0.08983226938927827, 0.7126622830878125], [-1.7036480556534283, 0.7960787201083437], [1.7035677028481488, -0.7961234727197022], [1.607106093246473, 0.5686524941018596], [-1.607102046898864, -0.568650772274404]]
The local minima for the Six-Hump Camel simulation function as evaluated by APOSMM with libEnsemble should be listed directly above.
Please see the API reference here for more APOSMM configuration options and other information.
Each of these example files can be found in the repository in examples/tutorials/aposmm.
Applications¶
APOSMM is not limited to evaluating minima from pure Python simulation functions. Many common libEnsemble use-cases involve using libEnsemble’s MPI Executor to launch user applications with parameters requested by APOSMM, then evaluate their output using APOSMM, and repeat until minima are identified. A currently supported example can be found in libEnsemble’s WarpX Scaling Test.